MELANOMA
Compiled by the APMG Molecular Committee- NORMAL MOLECULAR PATHWAYS AND ASSOCIATED MUTATIONS
- TEST COMMENTS
- TEST INTERPRETATION
- SUGGESTED READING
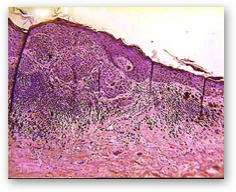
NORMAL MOLECULAR PATHWAYS AND ASSOCIATED MUTATIONS
The RAS-RAF-MEK-ERK signaling pathway (MAPK pathway) is a classical intracellular pathway that plays a crucial role in the homeostasis of normal cell turnover, cellular proliferation, differentiation, survival, and apoptosis. When activated aberrantly, this signaling pathway can induce tumorigenesis and has been associated with various malignancies.
MAPK Signaling Pathway

BRAF
Approximately 40-60% of melanomas and 70-80% of acquired melanocytic nevi have shown to harbor characteristic genetic alterations, most commonly activating mutations of BRAF at codon 600. BRAF is a protein kinase, which is a potent activator of MAPK pathway. It is located downstream of both EGFR and KRAS in the MAPK signaling pathway. Activating mutations of BRAF may lead to uncontrolled cellular proliferation and contribute to carcinogenesis.
BRAF V600 mutation analysis is often utilized for guiding treatment of patients with melanoma. The widely used commercially available methods for BRAF mutation detection in melanoma patients are allele-specific polymerase chain reaction (PCR) and bidirectional direct fluorescent sequencing.
If patients carry a BRAF V600 mutation, they may benefit from selective BRAF inhibitor treatment regimens. Various tyrosine kinase inhibitors of BRAF are available, including sorafenib, vemurafenib, PLX4032, and more. These targeted therapies show promising results in mutation-positive melanoma patients.
The most common and most routinely tested BRAF V600 mutation is V600E, which accounts for approximately 70 to 90% of BRAF mutated melanoma cases. V600K is the second most common mutation, accounting for approximately 6 to 29% of BRAF mutated melanoma cases. Other less common mutations include BRAF V600R and V600D. Each of the above mutations correspond to a different amino acid substitution involving codon 600.
Studies have shown that patients with BRAF V600K mutations may benefit from BRAF-inhibitor therapies, similar to patients harboring V600E mutations. Preclinical trials show that patients with BRAF V600D-mutated melanomas also show favorable response to BRAF-inhibitor therapies. Therefore, besides BRAF V600E (which is detected in 50% of melanomas), the addition of testing for mutations V600K and V600D, can help to identify another 10 to 15% patients, who could possibly benefit from gene-targeted therapies.
NRAS
Approximately 15-30% of cutaneous melanomas have mutations in the NRAS gene, which leads to a constantly active NRAS protein. Like BRAF mutations, NRAS mutations are driver mutations; melanomas with NRAS mutations may be dependent on NRAS activation for continued cellular proliferation and survival.
In comparison with melanomas with BRAF mutations, or melanomas with wild-type BRAF and NRAS, melanomas with NRAS mutations tend to be more aggressive clinically and histologically are associated with thicker tumors and higher mitotic rates. The prognosis is typically worse for tumors with NRAS mutation when compared to tumors with BRAF mutations, while tumors that are wild type for BRAF and NRAS tend to have the best prognosis.
NRAS and BRAF mutations in melanoma are usually mutually exclusive, especially in patients who have not been treated with any BRAF-targeted therapies. However, approximately 20% of patients that have clinical progression of their disease following BRAF-targeted therapy are found to also harbor NRAS mutations.
Testing for BRAF mutation in patients with metastatic melanoma is now routine practice, since the US Food and Drug Administration has approved BRAF-targeted therapies. Some clinicians choose to test for both BRAF and NRAS prior to initiation of treatment for metastatic melanoma. Other clinicians may only test for NRAS mutations if the melanoma was found to be BRAF wild-type, since the two mutations are usually mutually exclusive and NRAS-inhibitor treatments are still in clinical trials.
Continued research on NRAS-targeted therapy for NRAS-mutated melanomas has shown promising results. The most successful therapeutic strategy includes inhibiting downstream signaling of NRAS in MAPK pathway. Other potential therapies include reducing the expression of NRAS, targeting the membrane localization of NRAS, and using combination therapies involving the phosphatidylinositide 3-kinase (Pl3K) pathway. Most of the NRAS-targeted therapies are still in the development stage.
The presence of BRAF/NRAS mutations in melanocytic nevi have not been proven to be associated with an increased risk of malignant transformation into melanoma. Melanomas associated with nevi show a similar frequency of BRAF and NRAS mutations when compared to cutaneous melanomas without associated nevi. Nevis that harbor BRAF or NRAS mutations have not been found to have a higher chance to be associated with melanomas than wild type nevi. Therefore research does not suggest that BRAF or NRAS mutations contribute at a molecular level to transformation of melanocytic nevi into melanoma. Consequently, testing for BRAF or NRAS mutations in melanocytic nevi is generally not performed.
TEST COMMENTS
BRAF mutation analysis by PCR is available upon request. This test is intended to detect patients with melanocytic neoplasms that are positive for the BRAF V600 mutations (E,K,D). Patients with V600 mutations may respond to selective BRAF inhibitor therapies.
NRAS mutation analysis by PCR is also available upon request.
TEST INTERPRETATION
PCR GENE MUTATION ANALYSIS INTERPRETATION:
Positive = BRAF V600 (E, K, or D) Mutation Detected
Negative = BRAF V600 (E, K, or D) Mutation Not Detected
Positive = NRAS Mutation Detected
Negative = NRAS Mutation Not Detected
SUGGESTED READING
PCR GENE MUTATION ANALYSIS INTERPRETATION:
Sullivan RJ, Flaherty KT. BRAF in melanoma: pathogenesis, diagnosis, inhibition, and resistance. Journal of Skin Cancer: 2011.
http://www.hindawi.com/journals/jsc/2011/423239/
Kelleher FC, McArthur GA. Testing NRAS in melanoma. Cancer Journal: 2012.
Tschandl P, Berghoff AS, Preusser M, et al. (2013) NRAS and BRAF Mutations in melanoma-associated nevi and uninvolved nevi. PLoS ONE 8(7):e69639.
Ascierto PA, Kirkwood JM, Grob J, et al. The role of BRAF V600 mutation in melanoma. Journal of Translational Medicine: 2012.

